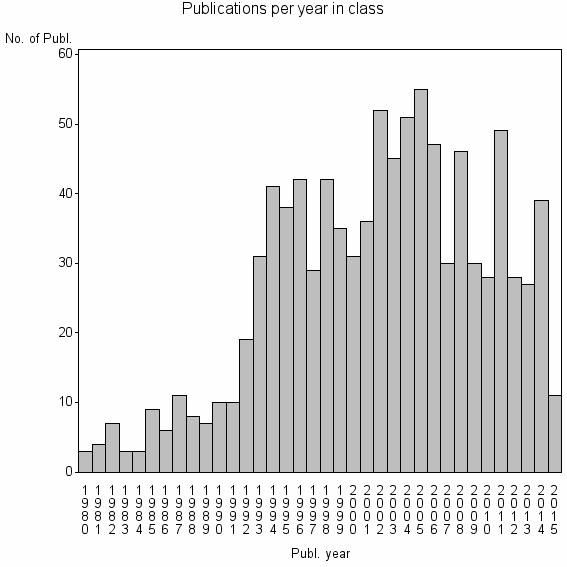
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 10834 | 963 | 29.3 | 64% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 2449 | 3598 | PREDICTION WITH EXPERT ADVICE//UNIVERSAL CODING//GRAPHICAL REPRESENTATION |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | LATENT PERIODICITY | Author keyword | 15 | 77% | 1% | 10 |
| 2 | 3 BASE PERIODICITY | Author keyword | 9 | 83% | 1% | 5 |
| 3 | DNA WALK | Author keyword | 9 | 67% | 1% | 8 |
| 4 | TRIPLET PERIODICITY | Author keyword | 8 | 75% | 1% | 6 |
| 5 | LONG RANGE CORRELATIONS | Author keyword | 6 | 18% | 3% | 30 |
| 6 | CODING NON CODING DNA SEQUENCES | Author keyword | 6 | 100% | 0% | 4 |
| 7 | A YA USIKOV RADIOPHYS ELECT | Address | 3 | 17% | 2% | 19 |
| 8 | DNA PERIODICITIES | Author keyword | 3 | 100% | 0% | 3 |
| 9 | THREE BASE PERIODICITY | Author keyword | 3 | 100% | 0% | 3 |
| 10 | INFORMATION DECOMPOSITION | Author keyword | 3 | 40% | 1% | 6 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | LATENT PERIODICITY | 15 | 77% | 1% | 10 | Search LATENT+PERIODICITY | Search LATENT+PERIODICITY |
| 2 | 3 BASE PERIODICITY | 9 | 83% | 1% | 5 | Search 3+BASE+PERIODICITY | Search 3+BASE+PERIODICITY |
| 3 | DNA WALK | 9 | 67% | 1% | 8 | Search DNA+WALK | Search DNA+WALK |
| 4 | TRIPLET PERIODICITY | 8 | 75% | 1% | 6 | Search TRIPLET+PERIODICITY | Search TRIPLET+PERIODICITY |
| 5 | LONG RANGE CORRELATIONS | 6 | 18% | 3% | 30 | Search LONG+RANGE+CORRELATIONS | Search LONG+RANGE+CORRELATIONS |
| 6 | CODING NON CODING DNA SEQUENCES | 6 | 100% | 0% | 4 | Search CODING+NON+CODING+DNA+SEQUENCES | Search CODING+NON+CODING+DNA+SEQUENCES |
| 7 | DNA PERIODICITIES | 3 | 100% | 0% | 3 | Search DNA+PERIODICITIES | Search DNA+PERIODICITIES |
| 8 | THREE BASE PERIODICITY | 3 | 100% | 0% | 3 | Search THREE+BASE+PERIODICITY | Search THREE+BASE+PERIODICITY |
| 9 | INFORMATION DECOMPOSITION | 3 | 40% | 1% | 6 | Search INFORMATION+DECOMPOSITION | Search INFORMATION+DECOMPOSITION |
| 10 | PROTEIN CODING REGION | 3 | 35% | 1% | 6 | Search PROTEIN+CODING+REGION | Search PROTEIN+CODING+REGION |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | 1 F ALPHA SPECTRUM | 61 | 95% | 2% | 20 |
| 2 | NONCODING DNA SEQUENCES | 39 | 61% | 4% | 42 |
| 3 | LONG RANGE CORRELATIONS | 39 | 20% | 18% | 172 |
| 4 | SPATIAL 1 F SPECTRA | 21 | 78% | 1% | 14 |
| 5 | FRACTAL CORRELATIONS | 21 | 73% | 2% | 16 |
| 6 | LINGUISTIC FEATURES | 18 | 55% | 2% | 22 |
| 7 | RANGE FRACTAL CORRELATIONS | 17 | 72% | 1% | 13 |
| 8 | NONCODING DNA | 15 | 25% | 6% | 55 |
| 9 | PROBABLE GENES | 15 | 73% | 1% | 11 |
| 10 | HIDDEN PERIODICITIES | 12 | 86% | 1% | 6 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Gene Prediction Based on DNA Spectral Analysis: A Literature Review | 2011 | 15 | 34 | 56% |
| Hierarchical structure of cascade of primary and secondary periodicities in Fourier power spectrum of alphoid higher order repeats | 2008 | 10 | 97 | 67% |
| The study of correlation structures of DNA sequences: a critical review | 1997 | 128 | 79 | 44% |
| Wavelets in bioinformatics and computational biology: state of art and perspectives | 2003 | 88 | 44 | 18% |
| Order and correlations in genomic DNA sequences. The spectral approach | 2000 | 16 | 83 | 61% |
| Wavelet analysis of DNA bending profiles reveals structural constraints on the evolution of genomic sequences | 2004 | 14 | 124 | 36% |
| Mathematical properties of DNA sequences from coding and noncoding regions | 2005 | 2 | 29 | 52% |
| Fractals from genomes | 2000 | 3 | 5 | 60% |
| Review of signal processing in genetics | 2005 | 3 | 69 | 30% |
| Gene identification in silico | 1996 | 1 | 32 | 41% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | A YA USIKOV RADIOPHYS ELECT | 3 | 17% | 2.0% | 19 |
| 2 | PHYS BIOL UNIT | 3 | 50% | 0.4% | 4 |
| 3 | BETH ISRAEL HOSP MED CARDIOVASC | 3 | 60% | 0.3% | 3 |
| 4 | CIELO | 2 | 67% | 0.2% | 2 |
| 5 | LINGUIST UNIT | 2 | 67% | 0.2% | 2 |
| 6 | PERSPECT INVEST | 2 | 30% | 0.6% | 6 |
| 7 | PROGRAM STAT OPERAT | 2 | 28% | 0.5% | 5 |
| 8 | BIOMOL PHYS MODELING GRP | 1 | 17% | 0.8% | 8 |
| 9 | ESCUELA INGN SISTEMAS COMPUTAC | 1 | 40% | 0.2% | 2 |
| 10 | GONDA GOLD MIED | 1 | 15% | 0.6% | 6 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000222544 | GRAPHICAL REPRESENTATION//ALIGNMENT FREE//SIMILARITIES DISSIMILARITIES |
| 2 | 0.0000205629 | COMPLEX QUANTITAT LINGUIST//MENZERATH ALTMANN LAW//FAMILY NAME DISTRIBUTION |
| 3 | 0.0000173678 | Q BIT//PAIR PREFERENCES//AMINO ACID DISTANCE MATRIX |
| 4 | 0.0000162558 | STELLA AVRAM GOREN GOLDSTEIN BIOTECHNOL EN//BACKBONE FRACTAL DIMENSION//BIOPHYS ENVIRONM PHYS |
| 5 | 0.0000148954 | GENE PREDICTION//GENE FINDING//GENE RECOGNITION |
| 6 | 0.0000122803 | DETRENDED FLUCTUATION ANALYSIS//BIOMED ELECT ENGN//MULTISCALE ENTROPY |
| 7 | 0.0000106393 | ELECTRON ION INTERACTION POTENTIAL//INFORMATIONAL SPECTRUM METHOD//DECEPTIVE IMPRINTING |
| 8 | 0.0000098597 | TRANSLATIONAL SELECTION//CODON USAGE//SYNONYMOUS CODON USAGE |
| 9 | 0.0000089680 | ISOCHORES//EVOLUZ MOL//ISOCHORE |
| 10 | 0.0000080998 | GENETIC CODE//EQUIPE BIOINFORMAT THEOR//CIRCULAR CODE |