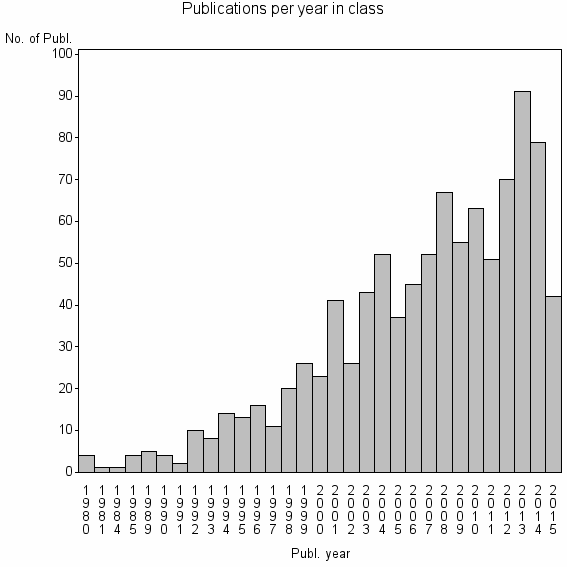
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 10675 | 976 | 50.7 | 91% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 195 | 20594 | PRE MRNA SPLICING//RNA LOCALIZATION//SPLICEOSOME |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | UPF1 | Author keyword | 66 | 79% | 4% | 42 |
| 2 | NMD | Author keyword | 58 | 53% | 8% | 77 |
| 3 | NONSENSE MEDIATED MRNA DECAY | Author keyword | 43 | 45% | 7% | 72 |
| 4 | EXON JUNCTION COMPLEX | Author keyword | 35 | 73% | 3% | 27 |
| 5 | NONSENSE MEDIATED DECAY | Author keyword | 21 | 38% | 5% | 45 |
| 6 | NONSENSE CODONS | Author keyword | 21 | 90% | 1% | 9 |
| 7 | Y14 | Author keyword | 19 | 76% | 1% | 13 |
| 8 | MAGOH | Author keyword | 17 | 100% | 1% | 8 |
| 9 | MRNA SURVEILLANCE | Author keyword | 16 | 54% | 2% | 20 |
| 10 | SMG 1 | Author keyword | 15 | 82% | 1% | 9 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | UPF1 | 66 | 79% | 4% | 42 | Search UPF1 | Search UPF1 |
| 2 | NMD | 58 | 53% | 8% | 77 | Search NMD | Search NMD |
| 3 | NONSENSE MEDIATED MRNA DECAY | 43 | 45% | 7% | 72 | Search NONSENSE+MEDIATED+MRNA+DECAY | Search NONSENSE+MEDIATED+MRNA+DECAY |
| 4 | EXON JUNCTION COMPLEX | 35 | 73% | 3% | 27 | Search EXON+JUNCTION+COMPLEX | Search EXON+JUNCTION+COMPLEX |
| 5 | NONSENSE MEDIATED DECAY | 21 | 38% | 5% | 45 | Search NONSENSE+MEDIATED+DECAY | Search NONSENSE+MEDIATED+DECAY |
| 6 | NONSENSE CODONS | 21 | 90% | 1% | 9 | Search NONSENSE+CODONS | Search NONSENSE+CODONS |
| 7 | Y14 | 19 | 76% | 1% | 13 | Search Y14 | Search Y14 |
| 8 | MAGOH | 17 | 100% | 1% | 8 | Search MAGOH | Search MAGOH |
| 9 | MRNA SURVEILLANCE | 16 | 54% | 2% | 20 | Search MRNA+SURVEILLANCE | Search MRNA+SURVEILLANCE |
| 10 | SMG 1 | 15 | 82% | 1% | 9 | Search SMG+1 | Search SMG+1 |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | EXON JUNCTION COMPLEX | 161 | 50% | 24% | 233 |
| 2 | SURVEILLANCE COMPLEX | 128 | 81% | 8% | 77 |
| 3 | NONSENSE MEDIATED DECAY | 81 | 34% | 20% | 195 |
| 4 | NMD | 79 | 79% | 5% | 50 |
| 5 | TRANSLATIONAL TERMINATION CODON | 75 | 76% | 5% | 52 |
| 6 | UPF1 | 67 | 77% | 5% | 46 |
| 7 | NMD FACTORS | 61 | 95% | 2% | 20 |
| 8 | CYTOPLASMIC TRANSLATION | 54 | 72% | 4% | 43 |
| 9 | UPF1 PROTEIN | 53 | 78% | 4% | 35 |
| 10 | UPF1 PHOSPHORYLATION | 45 | 83% | 3% | 25 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| NMD: a multifaceted response to premature translational termination | 2012 | 153 | 179 | 78% |
| Nonsense-mediated mRNA decay - Mechanisms of substrate mRNA recognition and degradation in mammalian cells | 2013 | 81 | 176 | 73% |
| The nonsense-mediated decay RNA surveillance pathway | 2007 | 539 | 161 | 81% |
| Organizing Principles of Mammalian Nonsense-Mediated mRNA Decay | 2013 | 55 | 192 | 65% |
| Nonsense-mediated mRNA decay in human cells: mechanistic insights, functions beyond quality control and the double-life of NMD factors | 2010 | 152 | 222 | 64% |
| Execution of nonsense-mediated mRNA decay: what defines a substrate? | 2009 | 135 | 102 | 84% |
| Nonsense-mediated mRNA decay: Splicing, translation and mRNP dynamics | 2004 | 685 | 156 | 63% |
| Nonsense-mediated mRNA decay: molecular insights and mechanistic variations across species | 2005 | 266 | 58 | 90% |
| Nonsense suppression therapies in ocular genetic diseases | 2015 | 1 | 48 | 77% |
| Quality control of eukaryotic mRNA: safeguarding cells from abnormal mRNA function | 2007 | 314 | 246 | 61% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | EDITH JOSEPH FI ER ENZYME INHIBITORS | 15 | 63% | 1.5% | 15 |
| 2 | MOL MED PARTNERSHIP UNIT | 7 | 28% | 2.3% | 22 |
| 3 | PEDIAT ONCOL HEMATOL IMMUNOL | 3 | 10% | 2.7% | 26 |
| 4 | MICROBIOL MOL BIODEF | 1 | 38% | 0.3% | 3 |
| 5 | EXPT THER EUT INTERDISCIPLINARY ONCOL | 1 | 100% | 0.2% | 2 |
| 6 | UMR IMAGING BRAIN | 1 | 100% | 0.2% | 2 |
| 7 | UMR8197 46 | 1 | 100% | 0.2% | 2 |
| 8 | ALBERT SHERMAN | 1 | 33% | 0.2% | 2 |
| 9 | BIOL SCI PROGRAM BK21 | 1 | 50% | 0.1% | 1 |
| 10 | CELL BIOLKIMMEL BIOL MED | 1 | 50% | 0.1% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000194580 | DECAPPING//P BODIES//CCR4 NOT |
| 2 | 0.0000145177 | MRNP BIOGENESIS METAB//DIS3//RRP6 |
| 3 | 0.0000096590 | SR PROTEINS//SR PROTEIN//ALTERNATIVE SPLICING |
| 4 | 0.0000093504 | FRAMESHIFTING//RIBOSOMAL FRAMESHIFTING//RNA PSEUDOKNOT |
| 5 | 0.0000088241 | RNA HELICASE//DEAD BOX//DEAD BOX PROTEIN |
| 6 | 0.0000074809 | TRANS TRANSLATION//TMRNA//SMPB |
| 7 | 0.0000074765 | INTRON GAIN//INTRON LOSS//INTRON EVOLUTION |
| 8 | 0.0000073650 | TRIGGER FACTOR//NASCENT CHAIN//COTRANSLATIONAL FOLDING |
| 9 | 0.0000063673 | SUP35//TRANSLATION TERMINATION//URE3 |
| 10 | 0.0000060665 | EXON SKIPPING//DMD GENET THER Y GRP//MC LOCKWOOD MUSCULAR DYSTROPHY |