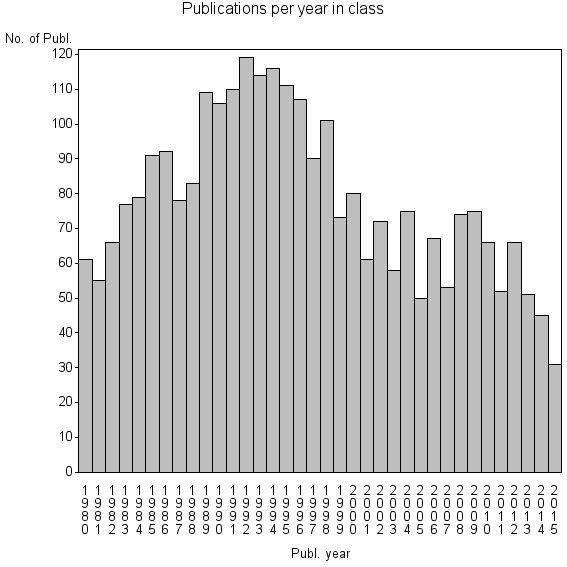
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 1009 | 2814 | 40.1 | 68% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | CYCLOSOPHORAOSES | Author keyword | 57 | 95% | 1% | 19 |
| 2 | LBMPS | Address | 39 | 73% | 1% | 30 |
| 3 | NOD GENES | Author keyword | 31 | 67% | 1% | 28 |
| 4 | NOD FACTOR | Author keyword | 23 | 45% | 1% | 38 |
| 5 | NOD FACTORS | Author keyword | 18 | 40% | 1% | 34 |
| 6 | NODULATION | Author keyword | 15 | 11% | 5% | 131 |
| 7 | NODD | Author keyword | 15 | 73% | 0% | 11 |
| 8 | RHIZOBIUM MELILOTI | Author keyword | 13 | 25% | 2% | 46 |
| 9 | NOD BOX | Author keyword | 13 | 80% | 0% | 8 |
| 10 | RHIZOBIUM | Author keyword | 13 | 10% | 4% | 118 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | CYCLOSOPHORAOSES | 57 | 95% | 1% | 19 | Search CYCLOSOPHORAOSES | Search CYCLOSOPHORAOSES |
| 2 | NOD GENES | 31 | 67% | 1% | 28 | Search NOD+GENES | Search NOD+GENES |
| 3 | NOD FACTOR | 23 | 45% | 1% | 38 | Search NOD+FACTOR | Search NOD+FACTOR |
| 4 | NOD FACTORS | 18 | 40% | 1% | 34 | Search NOD+FACTORS | Search NOD+FACTORS |
| 5 | NODULATION | 15 | 11% | 5% | 131 | Search NODULATION | Search NODULATION |
| 6 | NODD | 15 | 73% | 0% | 11 | Search NODD | Search NODD |
| 7 | RHIZOBIUM MELILOTI | 13 | 25% | 2% | 46 | Search RHIZOBIUM+MELILOTI | Search RHIZOBIUM+MELILOTI |
| 8 | NOD BOX | 13 | 80% | 0% | 8 | Search NOD+BOX | Search NOD+BOX |
| 9 | RHIZOBIUM | 13 | 10% | 4% | 118 | Search RHIZOBIUM | Search RHIZOBIUM |
| 10 | NODULATION GENES | 11 | 56% | 0% | 14 | Search NODULATION+GENES | Search NODULATION+GENES |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | CALCOFLUOR BINDING EXOPOLYSACCHARIDE | 201 | 86% | 4% | 103 |
| 2 | MELILOTI | 144 | 39% | 10% | 291 |
| 3 | NODULATION GENES | 138 | 58% | 6% | 158 |
| 4 | NODULE INVASION | 104 | 82% | 2% | 60 |
| 5 | RHIZOBIUM MELILOTI | 98 | 21% | 15% | 415 |
| 6 | CULTIVAR SPECIFIC NODULATION | 89 | 92% | 1% | 35 |
| 7 | FORM INEFFECTIVE NODULES | 83 | 84% | 2% | 46 |
| 8 | LEGUMINOSARUM | 83 | 35% | 7% | 191 |
| 9 | SP STRAIN NGR234 | 79 | 72% | 2% | 62 |
| 10 | LIPO OLIGOSACCHARIDE SIGNALS | 71 | 49% | 4% | 105 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Rhizobium lipo-chitooligosaccharide nodulation factors: Signaling molecules mediating recognition and morphogenesis | 1996 | 427 | 165 | 71% |
| Molecular basis of symbiotic promiscuity | 2000 | 371 | 228 | 83% |
| Signal molecules and cell-surface components involved in early stages of the legume-rhizobium interactions | 2015 | 2 | 259 | 58% |
| Surface polysaccharide involvement in establishing the rhizobium-legume symbiosis | 2003 | 160 | 116 | 96% |
| The roles of extracellular proteins, polysaccharides and signals in the interactions of rhizobia with legume roots | 2010 | 85 | 210 | 50% |
| Symbiotic use of pathogenic strategies: rhizobial protein secretion systems | 2009 | 80 | 98 | 64% |
| Environmental Signals and Regulatory Pathways That Influence Exopolysaccharide Production in Rhizobia | 2011 | 24 | 173 | 85% |
| THE RHIZOBIUM-PLANT SYMBIOSIS | 1995 | 291 | 235 | 80% |
| RHIZOBIUM - PLANT SIGNAL EXCHANGE | 1992 | 426 | 75 | 88% |
| Root nodulation and infection factors produced by rhizobial bacteria | 2000 | 220 | 161 | 85% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | LBMPS | 39 | 73% | 1.1% | 30 |
| 2 | BIOTECHNOL UBITA | 6 | 48% | 0.4% | 10 |
| 3 | BIOL MOL PLANTES SUPER | 4 | 38% | 0.3% | 9 |
| 4 | FB BIOL MOL ZELLBIOL ANGEW BOT | 3 | 100% | 0.1% | 3 |
| 5 | MOLEC PLANT SCI | 3 | 100% | 0.1% | 3 |
| 6 | UBIQUITOUS INFORMAT TECHNOL PLICAT CBRU | 3 | 30% | 0.3% | 9 |
| 7 | PHARMACOL TOXICOL FONDAMENTALES | 2 | 26% | 0.3% | 8 |
| 8 | BIO MOL INFORMAT BMIC | 2 | 67% | 0.1% | 2 |
| 9 | UMR 215 | 2 | 22% | 0.3% | 8 |
| 10 | INVEST TECNOL INNOVAC CITIUS | 2 | 36% | 0.1% | 4 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000196690 | SUPERNODULATION//LOTUS JAPONICUS//AUTOREGULATION OF NODULATION |
| 2 | 0.0000167469 | RHIZOBIA//BRADYRHIZOBIUM//RHIZOBIUM |
| 3 | 0.0000148970 | INOSITOL DEHYDROGENASE//INOSITOL CATABOLISM//MYO INOSITOL DEHYDROGENASE |
| 4 | 0.0000099076 | FDN PLANT NUTR//INDOLE 3 PYRUVIC ACID DECARBOXYLASE//L TRYPTOPHAN |
| 5 | 0.0000070723 | CU NUTRITION//COPPER NUTRITION//FERRICHROME UPTAKE |
| 6 | 0.0000067252 | PERIBACTEROID MEMBRANE//UREIDES//BACTEROIDS |
| 7 | 0.0000066848 | NIFA PROTEIN//SIGMA54//NIFA |
| 8 | 0.0000051870 | CROWN GALL//AGROBACTERIUM VITIS//TI PLASMID |
| 9 | 0.0000049327 | VITAVAX//BRADYRHIZOBIUM INOCULATION//CHLOROPYRIPHOS |
| 10 | 0.0000049118 | HEME SENSOR//FIXL//COOA |